DNA polymerase subunit gamma - Our Protein
DNA polymerase gamma is a type of DNA polymerase. DNA polymerases as a whole are enzymes that replicate DNA by attatching nucleotides which are essentially the building blocks of DNA that are either Adenine, Tyrosine, Cytosine our Guanine. These nucleotides are built in pairs that create a double stranded piece of DNA. The process of DNA replication is just copying a double strand of DNA into another double stranded piece of DNA. This is done via the DNA polymerase which "reads" the nucleotides and builds another copy of that strand. Cells need this process every time a cell divides. This is needed because otherwise we wouldn't be able to pass along the same genetic information from one cell to the next.
DNA polymerase gamma is a variation of DNA polymerase whereas DNA polymerase is often found in the nucleus of a cell, DNA polymerase gamma is found in the mitochondria of the cell. Here it replicates mitochondrial DNA (mtDNA) instead of the cellular DNA. mtDNA is different than the DNA found in the nucleus of the exact same cell. This is due because mtDNA function is used in the mitochondria and not the other parts of the cell. DNA polymerase gamma participates in proofreading (making sure mistakes during replication don't occur), repairing (repairs mistakes made during replication) and of course replication.
DNA polymerase gamma is a variation of DNA polymerase whereas DNA polymerase is often found in the nucleus of a cell, DNA polymerase gamma is found in the mitochondria of the cell. Here it replicates mitochondrial DNA (mtDNA) instead of the cellular DNA. mtDNA is different than the DNA found in the nucleus of the exact same cell. This is due because mtDNA function is used in the mitochondria and not the other parts of the cell. DNA polymerase gamma participates in proofreading (making sure mistakes during replication don't occur), repairing (repairs mistakes made during replication) and of course replication.
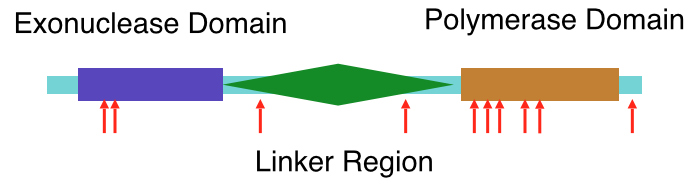
*SMART Diagram of the Human Protein DNA polymerase subunit gamma
*PFAM Diagram of the Human Protein DNA polymerase subunit gamma
Homology
Homology is essentially the similarity in characteristics due to the relatedness of organisms. Homologies can be found by looking at the anatomy of different organisms. Homology also can include looking at cellular similarities and differences as well as studying the vestigal structures that are found in organisms.
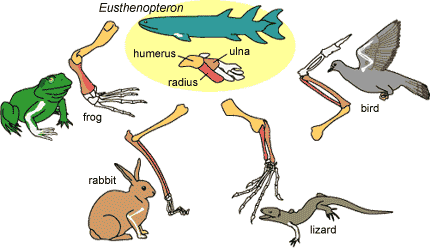
An example of homology is the forelimb of vertebrates with legs:
*photo credit
You can see that all of the organisms shown have different forelimbs, but they all share the same bones that make up the forelimb ;the humerus, the radius, and the ulna. These are the same bones found in their common ancestor, Eusthenopteron.
An example of homology is the forelimb of vertebrates with legs:
*photo credit
You can see that all of the organisms shown have different forelimbs, but they all share the same bones that make up the forelimb ;the humerus, the radius, and the ulna. These are the same bones found in their common ancestor, Eusthenopteron.
Homolog Reference
|
Human (Homo sapien) - DNA polymerase subunit gamma-1
Ascension Number: NP_001119603.1 FASTA Mouse (Mus Musculus) - DNA polymerase subunit gamma-1 isoform X2 Ascension Number: XP_006540772 FASTA E Value: 0 Identification (%): 89 Gorilla (Gorilla Gorilla) - DNA polymerase subunit gamma-1 Ascension Number: XP_004056785 FASTA E Value: 0 Identification (%): 99 Dog (Canis lupus familiarus) - DNA polymerase subunit gamma-1 Ascension Number: XP_005618410 FASTA E Value: 0 Identification (%): 87 Bonobo (Pan Paniscus) - DNA polymerase subunit gamma-1 isoform X2 Ascension Number: XP_008972123 FASTA E Value: 0 Identification (%): 98 African Elephant (Loxodonta africanas) - DNA polymerase subunit gamma-1 isoform X2 Ascension Number: XP_010592136 FASTA E Value: 0 Identification (%): 88 |
Anole lizard (Anolis carolinensis) - DNA polymerase subunit gamma-1 isoform X2
Ascension Number: XP_008118381 FASTA E Value: 0 Identification(%): 70 Alpaca (Vicugna pacos) - DNA polymerase subunit gamma-1 Ascension Number: XP_006198613 FASTA E Value: 0 Identification(%): 90 Hedgehog (Erinaceus europaeus) - DNA polymerase subunit gamma-1 Ascension Number: XP_007527320 FASTA E Value: 0 Identification(%): 84 Marmoset (Callithrix jacchus) - DNA polymerase subunit gamma-1 Ascension Number: XP_002749184 FASTA E Value: 0 Identification(%): 94 Medaka (Oryzias latipes) - DNA polymerase subunit gamma-1-like Ascension Number: XP_004070139 FASTA E Value: 0 Identification (%): 67 |
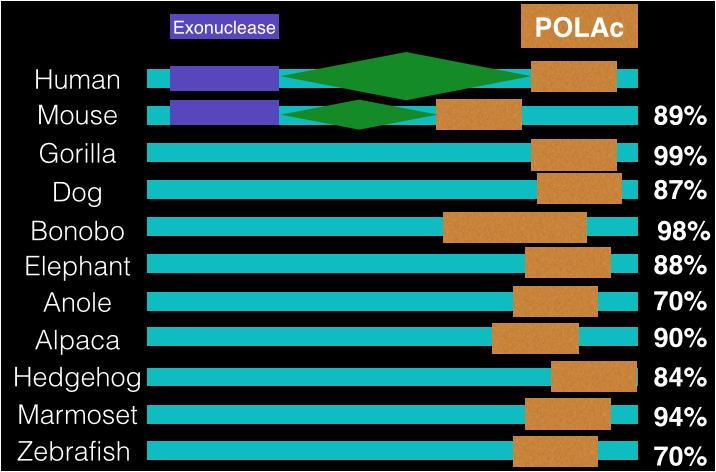
Protein Domains and Percent Identity
The protein motif above is comparing the POLG regions from 11 organisms. The region in orange is the coding region called POLAc. This region is believed to encode for DNA polymerase gamma. The green region found in humans and mice is the linker region and finally the purple region is the domain with exonuclease activity. You will notice that although the model organisms have high percent identity, that they lack the two domains found in mice and humans and this is due to the prediction software being unable to identify these region but experimentally they have been confirmed in both humans and mice. So there is a high possibility that these other animals have the same regions.
Analysis
From our homologs we can see that there is high level of similarity between the protein sequences. We can see in the animals with more developed brains that the sequences are much higher 99% and 98% similarity between the Gorilla and Bonobo respectively. All of the E Values are zero, which means that the significance of the similarity between the sequences is high. The lowest percentage identity is 67 is found in Medaka which is a fish and the farthest organism away in terms of species similarity. Due to the high identification percentage of these model organisms we can determine that the protein sequence in highly conserved. We can also tell from our protein motifs that the Protein coding region is highly conserved to include POLAc, which is the coding region for the DNA polymerase gamma. The sequences are relatively similar in the length of the coding region and similar in the position of the region. This helps confirm that this sequence is highly conserved.
References
- "Bioinformatics Lesson 3 -- Pairwise Alignment, Part 2." Bioinformatics Lesson 3 -- Pairwise Alignment, Part 2. VIBE Education Edition, n.d. Web. 25 Mar. 2015.
- "Domain Architecture Analysis." SMART: Sequence Analysis Results. N.p., n.d. Web. 25 Mar. 2015.
- "Family: DNA_pol_A (PF00476)." Pfam:. N.p., n.d. Web. 25 Mar. 2015
- Phenobank - Gene View (Y57A10A.15). N.p., n.d. Web. 24 Mar. 2015
- "Homologies." Understanding Evolution. UC Berkeley, n.d. Web. 25 Mar. 2015.